Animal Diversity
12.3 – Animal Phylogeny
Learning Objectives
By the end of this section, you will be able to do the following:
- Interpret the metazoan phylogenetic tree
- Describe the types of data that scientists use to construct and revise animal phylogeny
- List some of the relationships within the modern phylogenetic tree that have been discovered as a result of modern molecular data
Biologists strive to understand the evolutionary history and relationships of members of the animal kingdom, and all of life, for that matter. The study of phylogeny (the branching sequence of evolution) aims to determine the evolutionary relationships between phyla. Currently, most biologists divide the animal kingdom into 35 to 40 phyla. Scientists develop phylogenetic trees, which serve as hypotheses about which species have evolved from which ancestors.
Recall that until recently, only morphological characteristics and the fossil record were used to determine phylogenetic relationships among animals. Scientific understanding of the distinctions and hierarchies between anatomical characteristics provided much of this knowledge. Used alone, however, this information can be misleading. Morphological characteristics (such as skin color, body shape, etc.) may evolve multiple times, and independently, through evolutionary history. Analogous characteristics may appear similar between animals, but their underlying evolution may be very different. With the advancement of molecular technologies, modern phylogenetics is now informed by genetic and molecular analyses, in addition to traditional morphological and fossil data. With a growing understanding of genetics, the animal evolutionary tree has changed substantially and continues to change as new DNA and RNA analyses are performed on additional animal species.
Constructing an Animal Phylogenetic Tree
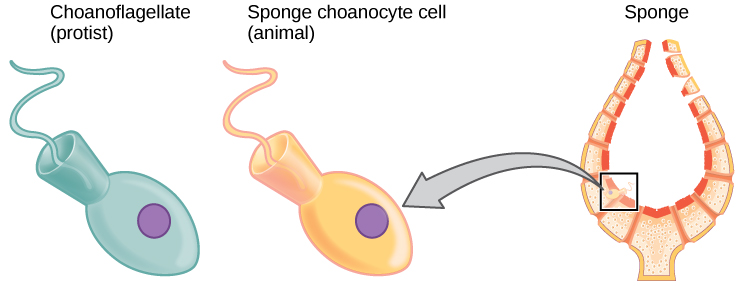
The current understanding of evolutionary relationships among animal, or Metazoa, phyla begins with the distinction between animals with true differentiated tissues, called Eumetazoa, and animal phyla that do not have true differentiated tissues, such as the sponges (Porifera) and the Placozoa. Similarities between the feeding cells of sponges (choanocytes) and choanoflagellate protists ((Figure)) have been used to suggest that Metazoa evolved from a common ancestral organism that resembled the moderncolonial choanoflagellates.
Eumetazoa are subdivided into radially symmetrical animals and bilaterally symmetrical animals, and are thus classified into the clades Bilateria and Radiata, respectively. As mentioned earlier, the cnidarians and ctenophores are animal phyla with true radial, biradial, or rotational symmetry. All other Eumetazoa are members of the Bilateria clade. The bilaterally symmetrical animals are further divided into deuterostomes (including chordates and echinoderms) and two distinct clades of protostomes (including ecdysozoans and lophotrochozoans) ((Figure)a,b). Ecdysozoa includes nematodes and arthropods; they are so named for a commonly found characteristic among the group: the physiological process of exoskeletal molting followed by the “stripping” of the outer cuticular layer, called ecdysis. Lophotrochozoa is named for two structural features, each common to certain phyla within the clade. Some lophotrochozoan phyla are characterized by a larval stage called trochophore larvae, and other phyla are characterized by the presence of a feeding structure called a lophophore (thus, the shorter term, “lopho-trocho-zoa”).

Link to Learning
Explore an interactive tree of life here. Zoom and click to learn more about the organisms and their evolutionary relationships.
Modern Advances in Phylogenetic Understanding Come from Molecular Analyses
The phylogenetic groupings are continually being debated and refined by evolutionary biologists. Each year, new evidence emerges that further alters the relationships described by a phylogenetic tree diagram.
Link to Learning
Watch the following video to learn how biologists use genetic data to determine relationships among organisms.
Nucleic acid and protein analyses have greatly modified and refined the modern phylogenetic animal tree. These data come from a variety of molecular sources, such as mitochondrial DNA, nuclear DNA, ribosomal RNA (rRNA), and certain cellular proteins. Many evolutionary relationships in the modern tree have only recently been determined from the molecular evidence. For example, a previously classified group of animals called lophophorates, which included brachiopods and bryozoans, were long-thought to be primitive deuterostomes. Extensive molecular analysis using rRNA data found these animals are actually protostomes, more closely related to annelids and mollusks. This discovery allowed for the distinction of the protostome clade Lophotrochozoa. Molecular data have also shed light on some differences within the lophotrochozoan group, and the placement of the Platyhelminthes is particularly problematic. Some scientists believe that the phyla Platyhelminthes and Rotifera should actually belong to their own clade of protostomes termed Platyzoa.
Molecular research similar to the discoveries that brought about the distinction of the lophotrochozoan clade has also revealed a dramatic rearrangement of the relationships between mollusks, annelids, arthropods, and nematodes, and as a result, a new ecdysozoan clade was formed. Due to morphological similarities in their segmented body types, annelids and arthropods were once thought to be closely related. However, molecular evidence has revealed that arthropods are actually more closely related to nematodes, now comprising the ecdysozoan clade, and annelids are more closely related to mollusks, brachiopods, and other phyla in the lophotrochozoan clade. These two clades now make up the protostomes.
Another change to former phylogenetic groupings because of modern molecular analyses includes the emergence of an entirely new phylum of worm called Acoelomorpha. These acoel flatworms were long thought to belong to the phylum Platyhelminthes because of their similar “flatworm” morphology. However, molecular analyses revealed this to be a false relationship and originally suggested that acoels represented living species of some of the earliest divergent bilaterians. More recent research into the acoelomorphs has called this hypothesis into question and suggested that the acoels are more closely related to deuterostomes. The placement of this new phylum remains disputed, but scientists agree that with sufficient molecular data, their true phylogeny will be determined.
Another example of phylogenetic reorganization involves the identification of the Ctenophora as the basal clade of the animal kingdom. Ctenophora, or comb jellies, were once considered to be a sister group of the Cnidaria, and the sponges (Porifera) were placed as the basal animal group, sister to other animals. The presence of nerve and muscle cells in both the Ctenophores and the Cnidaria and their absence in the Porifera strengthened this view of the relationships among simple animal forms. However, recent molecular analysis has shown that many of the genes that support neural development in other animals are absent from the Ctenophore genome. The muscle cells are restricted to the mouth and tentacles and are derived from cells in the mesoglea. The mitochondrial genome of the Ctenophores is small and lacks many genes found in other animal mitochondrial genomes. These features plus the absence of Hox genes from the Ctenophores have been used to argue that the Ctenophores should be considered basal or as a sister group of the Porifera, and that the evolution of specialized nerve and muscle tissue may have occurred more than once in the history of animal life. Although Ctenophores have been shown as basal to other animals in the phylogeny presented in Chapter 27.2, debate on this issue is likely to continue as Ctenophores are more closely studied.
Changes to the phylogenetic tree can be difficult to track and understand, and are evidence of the process of science. Data and analytical methods play a significant role in the development of phylogenies. For this reason – because molecular analysis and reanalysis are not complete — we cannot necessarily dismiss a former phylogenetic tree as inaccurate. A recent reanalysis of molecular evidence by an international group of evolutionary biologists refuted the proposition that comb jellies are the phylogenetically oldest extant metazoan group. The study, which relied on more sophisticated methods of analyzing the original genetic data, reaffirms the traditional view that the sponges were indeed the first phylum to diverge from the common ancestor of metazoans. The ongoing discussion concerning the location of sponges and comb jellies on the animal “family tree” is an example of what drives science forward.
Section Summary
Scientists are interested in the evolutionary history of animals and the evolutionary relationships among them. There are three main sources of data that scientists use to create phylogenetic evolutionary tree diagrams that illustrate such relationships: morphological information (which includes developmental morphologies), fossil record data, and, most recently, molecular data. The details of the modern phylogenetic tree change frequently as new data are gathered, and molecular data has recently contributed to many substantial modifications of the understanding of relationships between animal phyla.
Glossary
- Ecdysozoa
- clade of protostomes that exhibit exoskeletal molting (ecdysis)
- Eumetazoa
- group of animals with true differentiated tissues
- Lophotrochozoa
- clade of protostomes that exhibit a trochophore larvae stage or a lophophore feeding structure
- Metazoa
- group containing all animals
- Parazoa
- group of animals without true differentiated tissues

